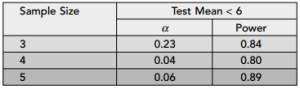
In his book Regression Modeling Strategies, 2nd Ed, Frank Harrell provides a list of what he calls the “most useful tests” for a 2-level factor \(\times\) numeric model (Table 2.2, p. 19). This is often called an Analysis of Covariance, or ANCOVA. The basic idea is we have a numeric response with substantial variability and we seek to understand the variability by modeling the mean of the response as a function of a categorical variable and a numeric variable.
Let’s simulate some data for such a model and then see how we can use R to carry out these tests.
n <- 400
set.seed(1)
sex <- factor(sample(x = c("f", "m"), size = n, replace = TRUE))
age <- round(runif(n = n, min = 18, max = 65))
y <- 1 + 0.8*age + 0.4*(sex == "m") - 0.7*age*(sex == "m") + rnorm(n, mean = 0, sd = 8)
dat <- data.frame(y, age, sex)
The data contain a numeric response, y, that is a function of age and sex. I set the “true” coefficient values to 1, 0.8, 0.4, and -0.7. They correspond to \(\beta_0\) through \(\beta_3\) in the following model:
\[y = \beta_0 + \beta_1 age + \beta_2 sex + \beta_3 age \times sex\]
In addition the error component is a Normal distribution with a standard deviation of 8.
Now let’s model the data and see how close we get to recovering the true parameter values.
mod <- lm(y ~ age * sex, dat)
summary(mod)
## ## Call: ## lm(formula = y ~ age * sex, data = dat) ## ## Residuals: ## Min 1Q Median 3Q Max ## -23.8986 -5.8552 -0.2503 6.0507 30.6188 ## ## Coefficients: ## Estimate Std. Error t value Pr(>|t|) ## (Intercept) 0.27268 1.93776 0.141 0.888 ## age 0.79781 0.04316 18.484 <2e-16 *** ## sexm 2.07143 2.84931 0.727 0.468 ## age:sexm -0.72702 0.06462 -11.251 <2e-16 *** ## --- ## Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1 ## ## Residual standard error: 8.661 on 396 degrees of freedom ## Multiple R-squared: 0.7874, Adjusted R-squared: 0.7858 ## F-statistic: 489 on 3 and 396 DF, p-value: < 2.2e-16
While the coefficient estimates for age and the age \(\times\) sex interaction are pretty close to the true values, the same cannot be said for the intercept and sex coefficients. The residual standard error of 8.661 is close to the true value of 8.
We can see in the summary output of the model that four hypothesis tests, one for each coefficient, are carried out for us. Each are testing if the coefficient is equal to 0. Of those four, only one qualifies as one of the most useful tests: the last one for age:sexm. This tests if the effect of age is independent of sex and vice versa. Stated two other ways, it tests if age and sex are additive, or if the age effect is the same for both sexes. To get a better understanding of what we’re testing, let’s plot the data with fitted age slopes for each sex.
library(ggplot2)
ggplot(dat, aes(x = age, y = y, color = sex)) +
geom_point() +
geom_smooth(method="lm")
Visually it appears the effect of age is not independent of sex. It seems more pronounced for females. Is this effect real or maybe due to chance? The hypothesis test in the summary output for age:sexm evaluates this. Obviously the effect seems very real. We are not likely to see such a difference in slopes this large if there truly was no difference. It does appear the effect of age is different for each sex. The estimate of -0.72 estimates the difference in slopes (or age effect) for the males and females.
The other three hypothesis tests are not very useful.
- Testing if the
Interceptis 0 is testing whetheryis 0 for females at age 0. - Testing if
ageis 0 is testing whetherageis associated withyfor males. - Testing if
sexmis 0 is testing whethersexis associated withyfor subjects at age 0.
Other more useful tests, as Harrell outlines in Table 2.2, are as follows:
- Is
ageassociated withy? - Is
sexassociated withy? - Are either
ageorsexassociated withy?
The last one is answered in the model output. That’s the F-statistic in the last line. It tests whether all coefficients (except the intercept) are equal to 0. The result of this test is conclusive. At least one of the coeffcients is not 0.
To test if age is associated with y, we need to test if both the age and age:sexm coefficents are equal to 0. The car package by John Fox provides a nice function for this purpose called linearHypothesis. It takes at least two arguments. The first is the fitted model object and the second is a vector of hypothesis tests. Below we specify we want to test if “age = 0” and “age:sexm = 0”
library(car)
linearHypothesis(mod, c("age = 0", "age:sexm = 0"))
## Linear hypothesis test ## ## Hypothesis: ## age = 0 ## age:sexm = 0 ## ## Model 1: restricted model ## Model 2: y ~ age * sex ## ## Res.Df RSS Df Sum of Sq F Pr(>F) ## 1 398 55494 ## 2 396 29704 2 25790 171.91 < 2.2e-16 *** ## --- ## Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
The result is once again conclusive. The p-value is virtually 0. It does indeed appear that age is associated with y.
Likewise, to test if sex is associated with y, we need to test if both the sex and age:sexm coefficents are equal to 0.
linearHypothesis(mod, c("sexm = 0", "age:sexm = 0"))
## Linear hypothesis test ## ## Hypothesis: ## sexm = 0 ## age:sexm = 0 ## ## Model 1: restricted model ## Model 2: y ~ age * sex ## ## Res.Df RSS Df Sum of Sq F Pr(>F) ## 1 398 119354 ## 2 396 29704 2 89651 597.6 < 2.2e-16 *** ## --- ## Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
As expected this test confirms that sex is associated with y, just as we specified when we simulated the data.
Now that we have established that age is associated with y, and that the association differs for each sex, what exactly is that association for each sex? In other words what are the slopes of the lines in our plot above?
We can sort of answer that with the model coefficients.
round(coef(mod),3)
## (Intercept) age sexm age:sexm ## 0.273 0.798 2.071 -0.727
That corresponds to the following model:
\[y = 0.273 + 0.799 age + 2.071 sex – 0.727 age \times sex\]
When sex is female, the fitted model is
\[y = 0.273 + 0.799 age \]
This says the slope of the age is about 0.8 when sex is female.
When sex is male, the fitted model is
\[y = (0.273 + 2.071) + (0.797 – 0.727) age \]
\[y = 2.344 + 0.07 age \]
This says the slope of the age is about 0.07 when sex is male.
How certain are we about these estimates? That’s what standard error is for. For the age slope estimate for females the standard error is provided in the model output for the age coefficient. It shows about 0.04. Adding and subtracting 2 \(\times\) 0.04 to the coefficient gives us a rough 95% confidence interval. Or we could just use the confint function:
confint(mod, parm = "age")
## 2.5 % 97.5 % ## age 0.7129564 0.8826672
The standard error of the age slope estimate for males takes a little more work. Another car function useful for this is the deltaMethod function. It takes at least three arguments: the model object, the quantity expressed as a character phrase that we wish to estimate a standard error for, and the names of the parameters. The function then calculates the standard error using the delta method. Here’s one way to do it for our model
deltaMethod(mod, "b1 + b3", parameterNames = paste0("b", 0:3))
## Estimate SE 2.5 % 97.5 % ## b1 + b3 0.07079277 0.04808754 -0.02345709 0.1650426
The standard error is similar in magnitude, but since our estimate is so small the resulting confidence interval overlaps 0. This tells us the effect of age on males is too small for our data to determine if the effect is positive or negative.
Another way to get the estimated age slopes for each sex, along with standard errors and confidence intervals, is to use the margins package. We use the margins function with our model object and specify that we want to estimate the marginal effect of age at each level of sex. (“marginal effect of age” is another way of saying the effect of age at each level of sex)
library(margins)
margins(mod, variables = "age", at = list(sex = c("f", "m")))
## Average marginal effects at specified values
## lm(formula = y ~ age * sex, data = dat)
## at(sex) age ## f 0.79781 ## m 0.07079
This does the formula work we did above. It plugs in sex and returns the estmimated slope coefficient for age. If we wrap the call in summary we get the standard errors and confidence intervals.
summary(margins(mod, variables = "age", at = list(sex = c("f", "m"))))
## factor sex AME SE z p lower upper ## age 1.0000 0.7978 0.0432 18.4841 0.0000 0.7132 0.8824 ## age 2.0000 0.0708 0.0481 1.4722 0.1410 -0.0235 0.1650